The Simons Center for Systems Biology: Finding Signals in the Noise

Discoveries in biology historically have been the province of experimentalists. But advances during the past decade in sequencing human and other genomes have brought forth a vast amount of data that has led to the nascent but burgeoning field called systems biology in which theoretical approaches to biology play a vital role. “Biology has been predominantly an experimental science,” said Arnold J. Levine, Professor in the School of Natural Sciences and head of The Simons Center for Systems Biology at the Institute for Advanced Study. “That is especially true of twentieth-century biology, which really did not have a strong theoretical component, but instead, was based on observations.”
Systems biology is a rapidly growing and increasingly important field. Under Professor Levine, The Simons Center is focused on research at the interface of molecular biology and the physical sciences. To move forward the frontiers of understanding of fundamental biological problems, Members and Visitors explore and mine large data sets of genomes of organisms, expression patterns of genes in normal and pathological conditions, the genetic diversity found in species, and clinical and molecular correlations. The Center draws researchers from an array of disciplines, including mathematics, physics, astrophysics, molecular biology, chemistry, and computer science.
“Their training in analytical methods enables our Members to separate the signals in large data sets from the noise of the data, to make statistical associations, to establish probabilities of events occurring, and to predict outcomes that are testable,” said Levine. “The theorist now plays a role in biology that did not exist before.”
In analyzing these large data sets of information, systems biologists are able to reconstruct virtual genomes based on probability and develop a better understanding of evolution, migration, and the probabilities of the distribution of genes in populations. “We can trace back through time the evolution of humans into different races, into different groups,” said Levine.
Current research involves using such data to look at systems at the molecular, cellular, and organismic levels, to understand how they develop and interact and to discern what goes wrong in certain disease states, as well as to address important problems in biology that concern genetics and genomics, polymorphisms and aspects of evolution, signal transduction pathways and networks, stress responses, and pharmacogenomics in cancer biology.
Before Levine arrived at the Institute, a program in theoretical biology had been initiated in 1998 with the five-year appointment of Dr. Martin Nowak. Nowak, now Professor of Mathematics and Biology at Harvard University and director of its Program for Evolutionary Dynamics, had, with Oxford University professor Sir Robert May, a former Member in the School of Natural Sciences (1971–72), developed a wide variety of mathematical models to address a broad range of problems in evolutionary biology and infectious diseases, from how infectious agents spread within individuals and the dynamics of viral resistance, to the mechanism of disease progression. At the Institute, Nowak led a small group of post doctoral scholars and visiting senior scientists whose theoretical research interests included the dynamics of infectious diseases, evolutionary genomics, genetic instability, tumor progression, the evolution of language, and evolutionary theory in general.
Professor Levine, a molecular biologist, joined the Institute as Visiting Professor in the School of Natural Sciences in 2002, assumed leadership of the Institute’sbiology initiative in July 2003, and subsequently established the Center for Systems Biology. Levine is a former President of The Rockefeller University (1998–2002) and professor (1984–98) and chair of the Department of Molecular Biology at Princeton University (1984–96). A widely acclaimed leader in cancer research, Professor Levine discovered in 1979 the p53 tumor suppressor protein, a molecule that inhibits tumor development and whose disruption is associated with an estimated 50 percent of human cancer. For this discovery, he has won numerous awards, including the Keio Medical Science Prize, from the Keio University Medical Science Fund, Japan, in 2000; the Alfred Knudson Award in Cancer Genetics from the National Cancer Institute in 2001; the Albany Medical Center Prize in Medicine and Biomedical Research in 2001; and a 2006 Freedom to Discover grant from Bristol-Myers Squibb.
When Professor Levine was appointed to the Faculty in July 2004, systems biology was formally made a program of the School of Natural Sciences. Last year the Center was renamed The Simons Center for Systems Biology to reflect the generosity of Trustee James H. Simons, a former Member in the School of Mathematics,and his wife Marilyn Hawrys Simons, who have endorsed the Center’s work with a $10 million challenge grant from their foundation, The Simons Foundation. The Institute will expand the systems biology program demonstrably this fall when it begins construction of anextension to Bloomberg Hall, which will house the Center and unite the School of Natural Sciences in one building.
Because of the Institute’s long history in mathematics, physics, and astrophysics, the program has attracted exceptionally talented physicists, chemists, mathematicians, computer scientists, and statisticians who see the potential of an interdisciplinary approach to biology. Harlan Robins, now an assistant professor in computational biology at the Fred Hutchinson Cancer Research Center in Seattle, was a trained physicist specializing in string theory when he arrived at the Institute as a Member in systems biology (2002–2006). “I didn’t know any biology when I got here, so I would say my entire career as a biologist developed here," said Robins. "The first two years, I put a considerable effort into developing a background in biology."
Raúl Rabadán, who spent the last htree years as a particle physics Member in the School of Natural Sciences, has moved to systems biology for a three-year membership, beginning this term. Nils Baas, who twice has been a Member in the School of Mathematics, will spend a six-month sabbatical as a joint mathematics and systems biology Member, beginning in January.
Although there are no laboratories at the Institute, the Center enjoys close interactions with the Cancer Institute of New Jersey; the Robert Wood Johnson Medical School; Lewis-Sigler Center for Integrative Genomics at Princeton University; BioMaPS Institute at Rutgers, The State University of New Jersey; IBM; Merck & Co.; and Bristol-Myers Squibb. "Many of the things done here are tested in laboratories," said Levine. "we try to make it a rule that everything we work on has a testable outcome."
Professor Levine has made it a priority to position the Center in the middle of the biological community, rather than to operate in isolation. "We hold symposia here to introduce the Institute to experimentalists," said Levine. The program also benefits from close collaborations with leading laboratory scientists who "jump right in and do experiments that either verify or contradict the theoretical work we do," Levine said. "In this sense then, biology is adopting the paradigm that physics starts with. The theorist can actually lead the experimentalist rather than the experimentalist leading the theorist."
During his time at the Center, Robins developed algorithms to identify various genome sequences and their implications. With fellow Member Michael Krasnitz, Robins designed an algorithm that uncovered hidden sequences that are underrepresentedin the human genome but are prominent in the HIV genome. “The idea is that these particular sequences cause HIV to be underexpressed in the human cell and, therefore, when people try to do things like make a DNA vaccine for HIV, they don’t get any product,” said Robins. “You may need to change these sequences in order to make a DNA vaccine.”
As described in this issue’s cover, Rabadán, Robins, and Professor Levine also analyzed a large quantity of genomic sequence data for human and avian versions of the Type A influenza virus. In studying this data, which spanned almost ninety years, the team was able to identify genomic features that differed between the human and avian viruses and observe their evolution. In particular, they were able to discern significant differences in sequence changes that occurred over time, suggesting the possible existence of a previously unknown, innate immune response in humans that has no counterpart in birds. They were also able to provide evidence supporting the hypothesis that the avian version of the virus entered the human population just prior to the 1918 pandemic, probably no earlier than 1910. “Now, one of the questions is: Is it possible for the virus to jump directly from birds into humans?” said Rabadán. “And the other is: If this is possible, how is it possible?”
The potential impact of systems biology on the future of biological research is enormous, and should affect everything from population genetics, where it can impart a tremendous amount of information about selection and the process of evolution, to environmental biology, where it can identify, for example, previously unknown organisms in the air and unidentified viruses in ocean water.
In the field of medicine, it may be possible to draw conclusions from genetic material that will guide treatment of diseases such as breast cancer, where genetic markers may be able to indicate which patients will and will not benefit from certain therapies. Levine proposes that advances in systems biology could eventually lead doctors to conduct routine genetic testing that would identify which diseases a patient is predisposed to—information a patient could then use to make lifestyle decisions to avoid developing those diseases. The pioneering research that will make such advances possible, said Levine, “has very important societal and economic value.”
Analyzing Type A Influenza Viruses
Type A influenza viruses are the only influenza viruses known to have caused human pandemics. The steady advance of one such virus, influenza A(H5N1), commonly known as avian flu, has brought with it fears of a new pandemic. To date, no confirmed human-to-human transmissions have been reported in A(H5N1), but it has become increasingly important to understand the possible host-virus interactions.
In a paper that will be published in a forthcoming issue of the Journal of Virology, Member Raúl Rabadán and former Member (2002-2006) Harlan Robins together with Arnold J. Levine, Professor in the School of Natural Sciences and head of The Simons Center for Systems Biology at the Institute, present the results of their research comparing genomic sequence data for human and avian versions of the Type A influenza virus.
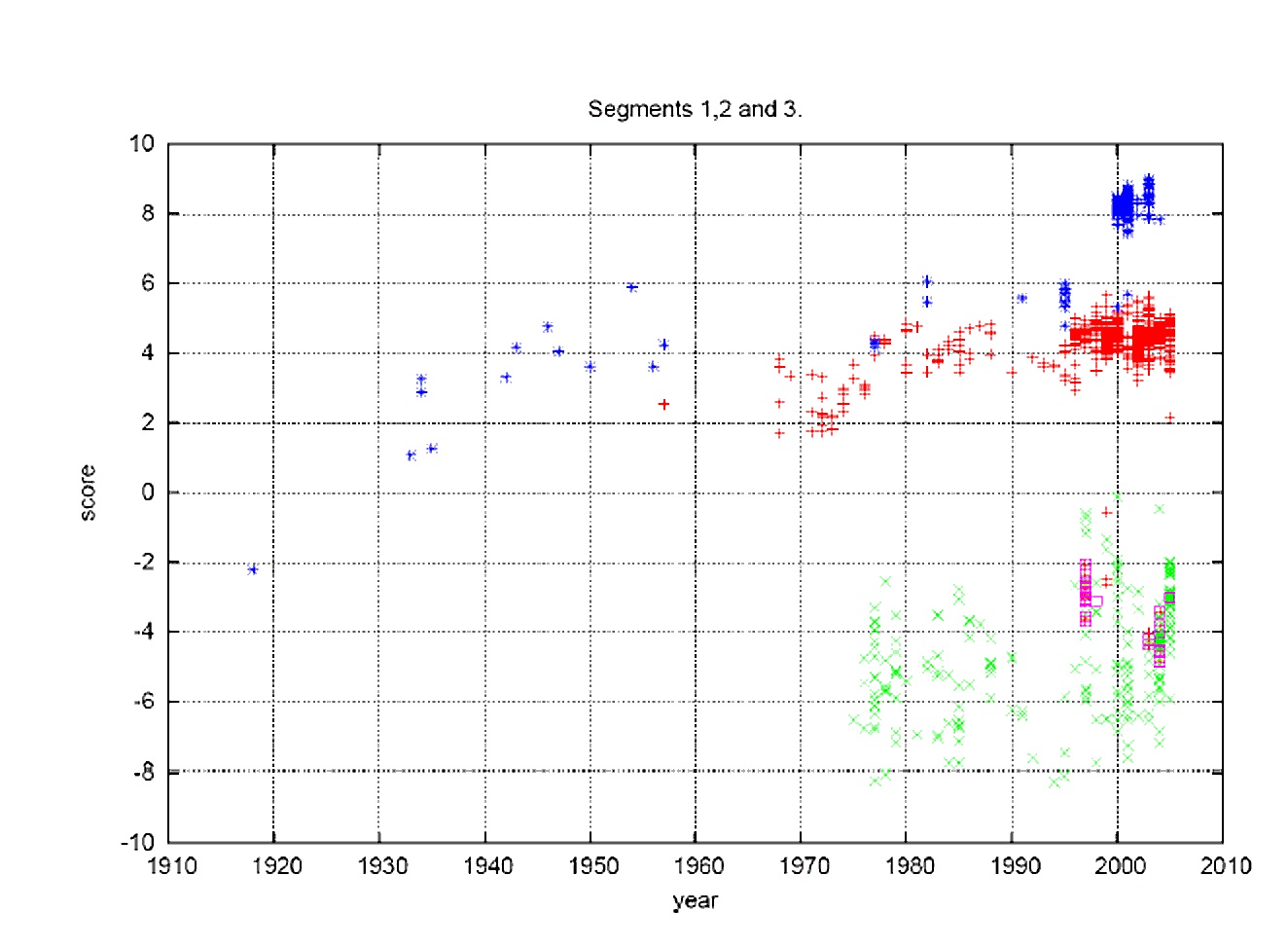
According to the authors, the genome of the influenza virus contains eight single-stranded negative RNA segments coding for ten or eleven proteins. When two or more different influenza viruses infect the same host cell, the new viral particles can incorporate replicas of segments from different original viruses. The graph above represents the added scores of the three longest RNA segments (1, 2 and 3). The x-axis shows the year when the influenza viruses were found: blue represents human H1N1 viruses; red shows human strains H2N2 (1957) and H3N2 (1968) as well as a few cases of H1N2 (US 2002) and H9N2 (Hong Kong 1999 of avian origin); green represents avian flu; and purple shows avian strains that jumped to humans but were not able to spread from human to human, including the present H5N1 strains. On the y-axis, viruses charted below zero are classified as avian and those charted above zero are classified as human.
As the graph demonstrates, the team was able to identify genomic features that distinguish the human and avian viruses and to observe their evolution. In studying the data, which spanned almost ninety years, they were able to discern significant differences in sequence changes that occurred over time, suggesting the possible existence of a previously unknown, innate immune response in humans that has no counterpart in birds. They were also able to provide evidence supporting the hypothesis that the avian version of the virus entered the human population (possibly after evolving in another organism for a period of time) just prior to the 1918 outbreak, probably no earlier than 1910.