Using Genetic Data to Revolutionize Understanding of Migration History
Few historical questions have so fascinated historians as the fall of the Roman Empire or, in the more fashionable modern parlance, its “transformation” into something altogether different, namely independent kingdoms ruled by successors of barbarian commanders in the West and a Greek-speaking Byzantine Empire in the East. For over two centuries, historians have particularly debated the role of barbarian invasions in this process, but in reality we have very little hard data on the nature of the barbarian “peoples” that entered the Western provinces between the fourth and sixth centuries, their numbers, their composition, or the reality of their influence on the indigenous populations of the Empire.
Were these large ethnic populations moving across Europe from Scandinavia to Italy and Spain, as nineteenth-century romantics imagined? Or were they small heterogeneous military units employed by the Empire that settled, with a minimum of force and disruption, within the administrative and fiscal mechanisms of a still-functioning Empire, as has been suggested more recently? Did these groups long maintain their distinctiveness from the local population, eschewing intermarriage and holding fast to their distinctive legal and cultural traditions, or did they rapidly integrate themselves into local elites through intermarriage and cultural transformation? Were they really distinct population groups at all or merely provincial Romans and local “barbarians” who united under ethnic labels and took advantage of opportunities to seize power from a beleaguered empire? Traditional sources with which to answer these questions—highly rhetorical accounts of the period often written centuries later, sparse administrative documents surviving in scattered fragments, and ambiguous archaeological material showing changing patterns of burial custom and settlements—simply do not provide enough evidence to reach a consensus.
More recently, a new kind of source, genetic data, has begun to be employed in an attempt to gain a new perspective on migration-era Europe, part of the widespread popularity of DNA research and a tendency to look to “science” to cut through the fuzzy speculations of historians and humanists. In an age obsessed with identity politics that looks increasingly to genetics to answer the fundamental question “Who am I?”, distributions of genetic markers across populations are being seen as proxies for ethnic and racial differences. In the words of Keith Wailoo, a historian of science and public affairs at Princeton University, the result is “lending renewed authority to biological conceptions of human difference and providing fodder for national debates over belonging, self-definition, and political power.”1 It is also producing bad history.
To cite but one example, a recent study announced the discovery of a “western Eurasian” in a two-thousand-year-old elite Xiongnu cemetery in northeast Mongolia and suggested that the presence of an “Indo-European” was possibly evidence of “the racial tolerance of the Xiongnu.”2 Can genetic analysis actually demonstrate that an individual was from a specific region of the Eurasian landmass, in this case from western Eurasia? Can it actually be evidence that he spoke an Indo-European language? Can it be evidence that the Xiongnu were racially tolerant? Such claims are wildly exaggerated and such
conclusions deeply problematic. What they actually found was an individual with the maternal U2e1 and paternal R1a1 haplogroups. R1a is the most common haplogroup in Europe, suggesting that it was statistically more likely but not at all certain that at some point his paternal ancestors had a western origin, and less likely that they were indigenous to the region in which he died. The fact that his genetic makeup bore some similarities with some modern Indian populations is absolutely no basis on which to term him an “Indo-European”—this is a linguistic, not a geographical, and certainly not a genetic term, and genotype is not equivalent to “race.” Such shifts from a biological to an essentializing cultural identity, common in the nineteenth and the first half of the twentieth centuries, have no place in modern scholarship. Other studies, based on DNA sampling of contemporary European populations, have attempted to argue for a massive replacement of male chromosomal material in eastern England between the fifth and eighth centuries, suggesting either the slaughter and expulsion of virtually all of the male British population of eastern Britain or else a form of “Anglo-Saxon apartheid” practiced by Anglo-Saxon invaders at the end of the Roman Empire.
Is this the best that genetics can offer migration history? I don’t think so, and I would like to suggest some possible areas in which genetic history might actually contribute to research projects exploring migration history. If, instead of using genetic evidence to look for essences, one uses this evidence to uncover one aspect of social construction, of transcultural and trans-societal flows, of movement and of transformation, then genetic history becomes both meaningful and exciting. The problem, of course, is how to do this.
Until recently, most attempts to use genetic evidence to write history suffered from two fundamental problems. First, until a decade or so ago, most historians attempted to work with contemporary DNA samples taken from living persons to trace their ancestors in order to understand earlier populations. While this may be effective for very early events (such as the dispersion of humans from Africa across Eurasia), it is very dubious evidence of more recent historical events such as medieval migrations, because it assumes extremely stable communities both before and after the events one hopes to study. Secondly, if scientists attempted to work with DNA taken from medieval graves, relatively little genetic data could be extracted from ancient tissue. This was essentially mtDNA, a nonrecombinant portion of one’s DNA inherited exclusively from one’s mother. Moreover, this data is such a small portion of the entire human genome that to base conclusions from it alone is, to paraphrase a metaphor of the British geneticist Mark Thomas, like reading a single page of a five-hundred-page book and drawing some conclusion about its contents. The result has been to ignore the complexity of the wider genetic data to essentialize communities by hiding the complexity of human ancestry.
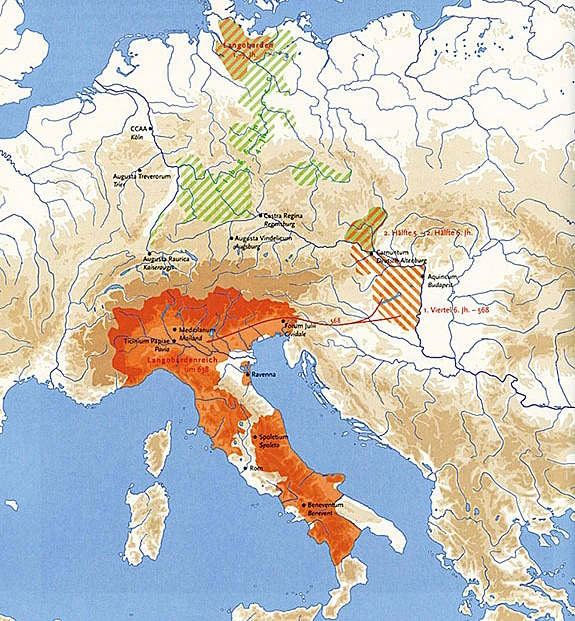
Today, thanks to extraordinary advances in genetic sequencing, it is at last possible to analyze the entire human genome of long-dead individuals and to study ancient populations at an extraordinarily fine-grained level. Using this “next-generation sequencing” it is possible to look at past populations in their complex genetic heterogeneity, to recognize fairly distant kinship relationships, and to gauge genetic distances separating individuals and populations. In the most ambitious attempt to apply these new techniques to study medieval migrations to date, I have assembled a team of geneticists, archaeologists, physical anthropologists, and historians to study migration of people living in Pannonia (what is now eastern Austria, Hungary, and the Czech Republic) into Italy in the second half of the sixth century. Historically, this migration/invasion is credited to the Longobardi, or Lombards, who established a kingdom in Italy that endured until the late eighth century and whose name lives on in Lombardia in Italy. Without assuming the veracity of historical sources written centuries later that tell of King Alboin leading his Longobards across the Alps in 568, we are studying the DNA of hundreds of individuals from cemeteries in Pannonia and Italy. Using next-generation sequencing, we hope to target distinct parts of the genome: approximately five thousand distinct locations along the genome called single nucleotide polymorphisms, or SNPs, are known to be useful in differentiating individuals from different regions of Europe and will allow us to look at close kinship among our populations; and five thousand regions of one thousand base pairs of continuous DNA sequence (a total of five megabases or 0.2 percent of the whole genome) will allow us to test competing models of the demographic history of this region. The results of this sequencing, which will take place in laboratories in Florence and Milan, must then be analyzed, both by examining what is termed unsupervised analysis—that is, direct analysis of the SNPs using such tools as principal component analysis and ancestry component analysis in order to find actual relationships and patterns—and, just as importantly, by testing a wide range of models derived from population genetics, cultural archaeology, and historical records, that might allow us to test the relative probability that any of the models might best explain our data. Ultimately, we hope to be able to construct a model of the populations of Pannonia and Italy in the late sixth century that can help us understand whether individuals identified by cultural archaeologists as “Longobards” based on their grave goods have closer genetic relationships with each other than with individuals buried according to what appear to be very different cultural traditions. We want to know if there are close relationships between similar cultural groups in Pannonia and Italy, or if the apparent cultural differences mask genetic homogeneity. We hope to determine whether men and women seem to have different histories of migration: do men move and marry local women, or are men more fixed and bring women from distant areas into their homes?
What we will not do is try to identify which among our samples are “real” Longobards or whether the Longobards are the “real” ancestors of the modern inhabitants of Lombardy. We want to get beyond ethnic and political labels and understand the movements of people and their cultural and demographic impacts on the population of Europe in the past. The project is at its initial phase of collecting genetic samples and doing the preliminary sequencing. We still face formidable obstacles: ancient DNA is notoriously difficult to sequence; obtaining good samples requires complex international negotiations and unusual interdisciplinary
cooperation; and the costs of next-generation sequencing and analysis are considerable. Nevertheless, I am energized by the opportunity to lead a pioneering project with the potential of revolutionizing our understanding of the population changes that transformed the Roman world at the end of antiquity.
1 Keith Wailoo, Alondra Nelson, Catherine Lee, eds., Genetics and the Unsettled Past: The Collision of DNA, Race, and History (Rutgers University Press, 2012), 2.
2 Kijeong Kim, et al., “A Western Eurasian Male is Found in 2000-Year-Old Elite Xiongnu Cemetery in Northeast Mongolia,” American Journal of Physical Anthropology 142, no. 3 (2010): 429–40.